A recap of CDSB 2018: the start of a community
Organizing an event is a lot of work, much more than you plan for it. But harvesting the fruits of a successful event makes the workload worth it.
In July, we kicked off the first Bioconductor Latin American Developers Workshop 2018 (LatAmBioc2018). The main goals of this meeting were to encourage the development of the Bioconductor project in Mexico and to start a community of Mexican bioinformatic software developers – the Community of Bioinformatic Software Developers (CDSB, from its acronym in Spanish).
The program of the workshop included talks, posters and lessons sessions as well as a two-day hands-on development hackathon. We had over 30 participants with strong quantitative background and experience in R programming from several LatAm countries.
Excited about #CDSBMexico? It starts tomorrow!
— ComunidadBioInfo (@CDSBMexico) July 29, 2018
¿Listos para mañana? 🇲🇽https://t.co/UUhX5RMSut
Schedulehttps://t.co/BZTD7wGskI
Materialshttps://t.co/Oz2ingGije#rstats #teaching @Bioconductor Thanks to @lcgunam @ccg_unam @LIIGH_UNAM
SoIBio @RConsortium@WINTERGENOMICS pic.twitter.com/YQxmnTNwQa
Community building
Unlike the US and Europe, where many universities have Computational Biology and Biostatistics Departments that gather experts in the field, in Mexico and Latin America experts in Biostatistics and Computational Biology are often spread across the country. The benefits of having Computational Biology and Biostatistics Departments are plenty: for example, experts can interact and work together on a specific problem, and researchers from other disciplines can easily find support and establish collaborations. The goal of CDSB is to create a virtual department where experts in Biostatistics and Computational Biology can discuss ideas and help each other across all stages of their careers.
The first event organized by CDSB, LatAmBioc2018 included talks where both participants and speakers could start a dialogue and motivate each other by showcasing their work. As keynote invited speakers we had Daniel Piñero, who talked about population genetics of maize in Mexico; Alejandro Ponce-Mendoza, who showed elegant shiny apps to visualize ecological data along with maps; Teresa Ortiz, who talked about gender biases among R programmers; and Benilton Carvalho, who showed the development of personalized medicine in Brazil.
We are having a great presentation from the founder of @RLadiesCDMX, @TeresaOM, about getting rid of gender biases among the #rstats community. #LatAmBioc18 @CDSBMexico #womeninSTEM pic.twitter.com/TPdDzHVdqC
— Alejandro Reyes (@areyesq) August 2, 2018
The LatAmBioc2018 poster session was designed to encourage interdisciplinary collaborations. We invited people to contribute bioinformatic problems that would benefit from advice from experts such as the participants of LatAmBioc2018. We were happy to see that this event helped to create new collaborations.
Here, we are presenting the basic structure and some examples with the framework -Winterflow- using @nextflowio, @Docker, #mk and #git pic.twitter.com/Ktn9o01X2Z
— Winter Genomics (@wintergenomics) August 2, 2018
A LatAm Bioconductor community
Anecdotally, some of us felt that Latin Americans were not fully represented in the Bioconductor Developers Meetings (see Leonardo Collado Torres’s blog post). As scientists, we like our statements to be supported by data: the map below shows the affiliations, when available, of the contributors of the latest package submissions to the Bioconductor project. (These data were scraped using the Github API.) Although biases such as Mexican contributors with affiliations abroad might not have been considered, it is clear that only a few contributions come from Latin American countries.
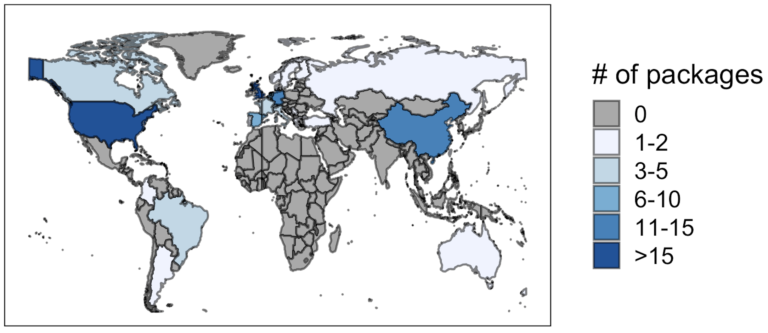
In hopes of filling this gap, we gathered a diverse team of instructors, including Martin Morgan, Benilton Carvalho and Selene L. Fernandez-Valverde, who taught participants how to develop R/Bioconductor packages. The main idea was to turn Bioconductor software users into Bioconductor software developers.
Let's estimate Pi! Rcpp exercise! Rcpp sugar! Now I'm hungry, I want pie @benilton @CDSBMexico #rstats #CDSBMexico #TIBMexico2018 🍰👩💻 pic.twitter.com/2OeGWr35r3
— Alejandra Medina-Rivera (@AleMedinaRivera) August 1, 2018
How to debug R code with Martin Morgan :D @CDSBMexico
— Semiramis (@semiramis_cj) August 1, 2018
Participants attended lectures from Monday to Wednesday. Before the workshop, co-organizer Daniela Ledezma received and coordinated R package development proposals. On Wednesday afternoon, the participants formed teams and chose problems from Daniela’s curated list. They worked on their projects on Thursday and Friday, and we were very pleased with the final products: most teams wrote documented R packages that are available through GitHub. Two interesting examples are the prototype of the regulonDB package, which enables users to query RegulonDB within an R interphase, and rGriffin, an R interphase to inquire regulatory networks (for details, check
their contributed blog post).
Participants are gathering to work in collaborative projects targeting specific questions. The objetive is write a fully documented #rstats package by Friday. @CDSBMexico #LatAmBioc18 pic.twitter.com/vDAvuOothC
— Alejandro Reyes (@areyesq) August 1, 2018
#CDSBMexico package developers course attendants presenting their projects #TIBs2018 #rstats @CDSBMexico pic.twitter.com/ST44hCsLT6
— Alejandra Medina-Rivera (@AleMedinaRivera) August 3, 2018
We are looking forward to seeing how the participants keep developing their R packages and collaborating through CSDB.
Overachievers
We felt that the goals of the workshop were reached programmers from all over Latin America worked together and wrote new R packages, which hopefully will be submitted to Bioconductor soon.
We also had the chance to enjoy some good Mexican food and hang out with colleagues in less formal settings, such as lunches and group outings. To celebrate the conclusion of the workshop, we had an improvised party in a bar that played Latin American music. As any good party should, it ended up quite late. Some of us went to bed at 4am, but we woke up energized to keep CSDB going. Even now, two months on, we are excited to continue developing a strong community of bioinformatic developers in Latin America.
In #mexico, we don't only know how to do #Science and good #Bioinformatics but also how to eat! Tacos during #TIBMexico2018 and @qBio18 @AlexielMedyna @licloona1487 something great is happening here 😊 pic.twitter.com/bC7ycfIA9n
— Ana B Villaseñor-Altamirano (@AnaBetty2304) August 2, 2018
Thank you
The LatAmBioc2018 would not have been possible without the time and support from the instructors, presenters, organizers and our sponsors CCG-UNAM, LCG-UNAM, LIIGH-UNAM, Winter Genomics, R Consortium and SoIBio.
